Suffix Trees

Overview
A suffix tree is a popular tool for analysing text strings. A Suffix Tree for a given text is a compressed trie that contains all of the text's suffixes. It's a type of digital tree that reveals the structure of a string and its subsets using algorithmic methods.
Takeaways
Applications of the Suffix tree
- Exact String Matching
- Computational Biology
- Text Statistics
- Longest Palindrome
- Data compression
What is Suffix Tree?
The prerequisite for this article is "trie". So firstly let's take a quick overview on trie.
Overview of Trie: In a trie data structure each alphabet of all the strings in the given string is traversed one after another one, and each node represents a single alphabet. The identical sub-string is represented by the same chain of nodes if two or more words begin with the same sub-string. Where the sub-string finishes and the unique suffix begins, the chain breaks. Each alphabet of the remaining suffix is then represented by its own node.
Now let's move on to the suffix tree for which we are here for.
Suffix tree is nothing but an extended version of trie. It's a compressed trie which includes all of a string's suffixes. There are some string-related problems which can be solved using suffix trees. Some of those problems are pattern matching, identifying unique substrings inside a string, and determining the longest palindrome. The suffix tree for the string S of length n is defined as a tree that has the following properties:
- There are exactly n leaves on the tree, numbered 1 through n.
- Each edge is identified by a non-empty S substring.
- Every internal node has at least two child, with the exception of the root.
- String-labels can't start with the same character on two edges that emerge from the same node.
- The suffix S[i..n], for I from 1 to n, is formed by combining all the string-labels encountered on the path from the root to the leaf i.
String S is padded with a terminal symbol not visible in the string, which is commonly marked $, because such a tree does not exist for all strings. There will be n leaf nodes, one for each of the n suffixes of S, and no suffix would be a prefix of another. Because all internal non-root nodes branch, there can only be n 1 such nodes, for a total of n + (n 1) + 1 = 2n nodes (n leaves, n 1 internal non-root nodes, and 1 root).
Example of Suffix tree
For making things more clear we will be first go through an example of trie which is given below.
There is list given below having name of some friends:
Below is a simplified trie representation for our list having friend names given below.
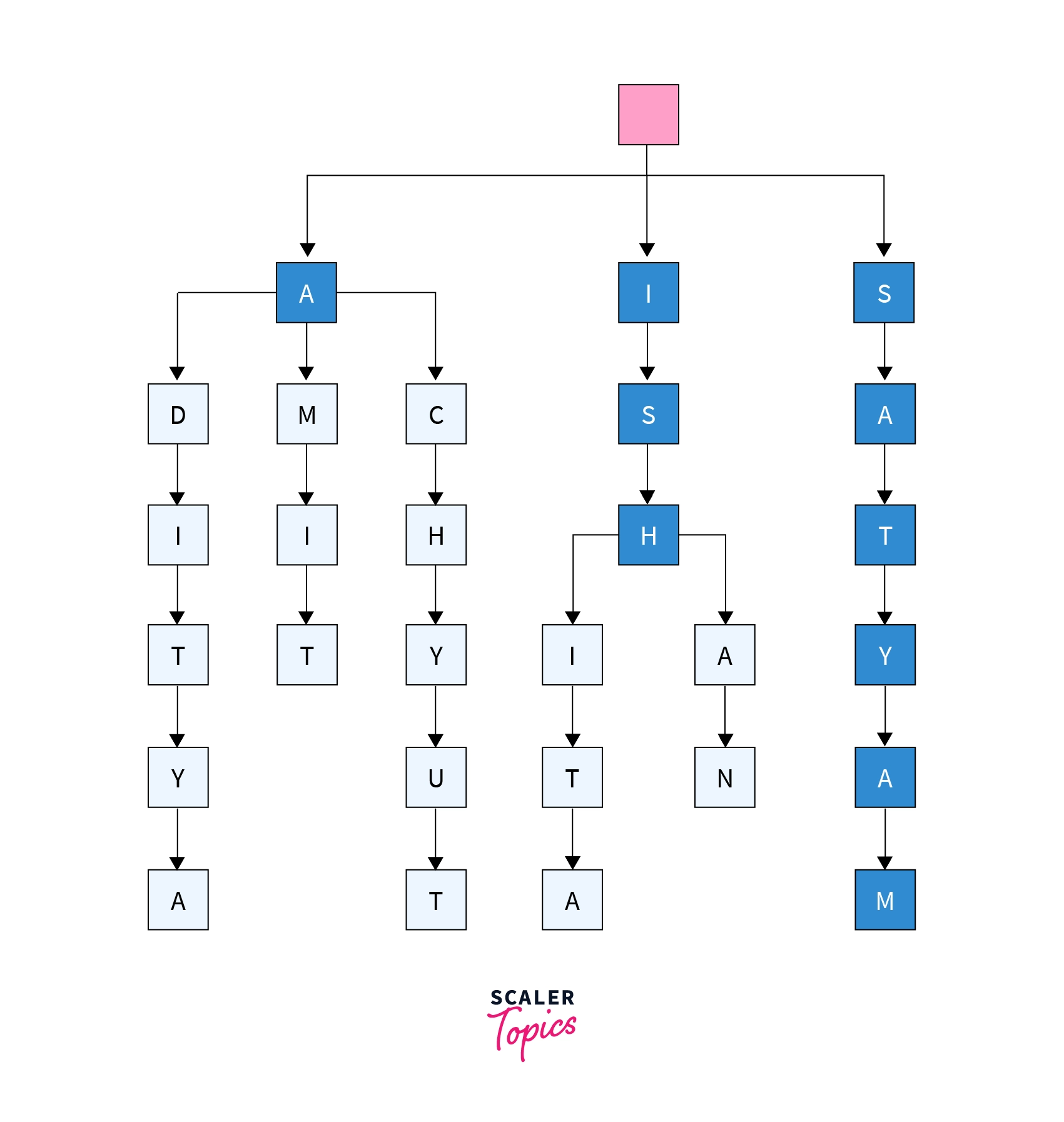
As previously stated, each distinct suffix in the list is compacted or compressed into a suffix tree. Below is a representation of the suffix tree for the same list of friend names that we used earlier for making trie.
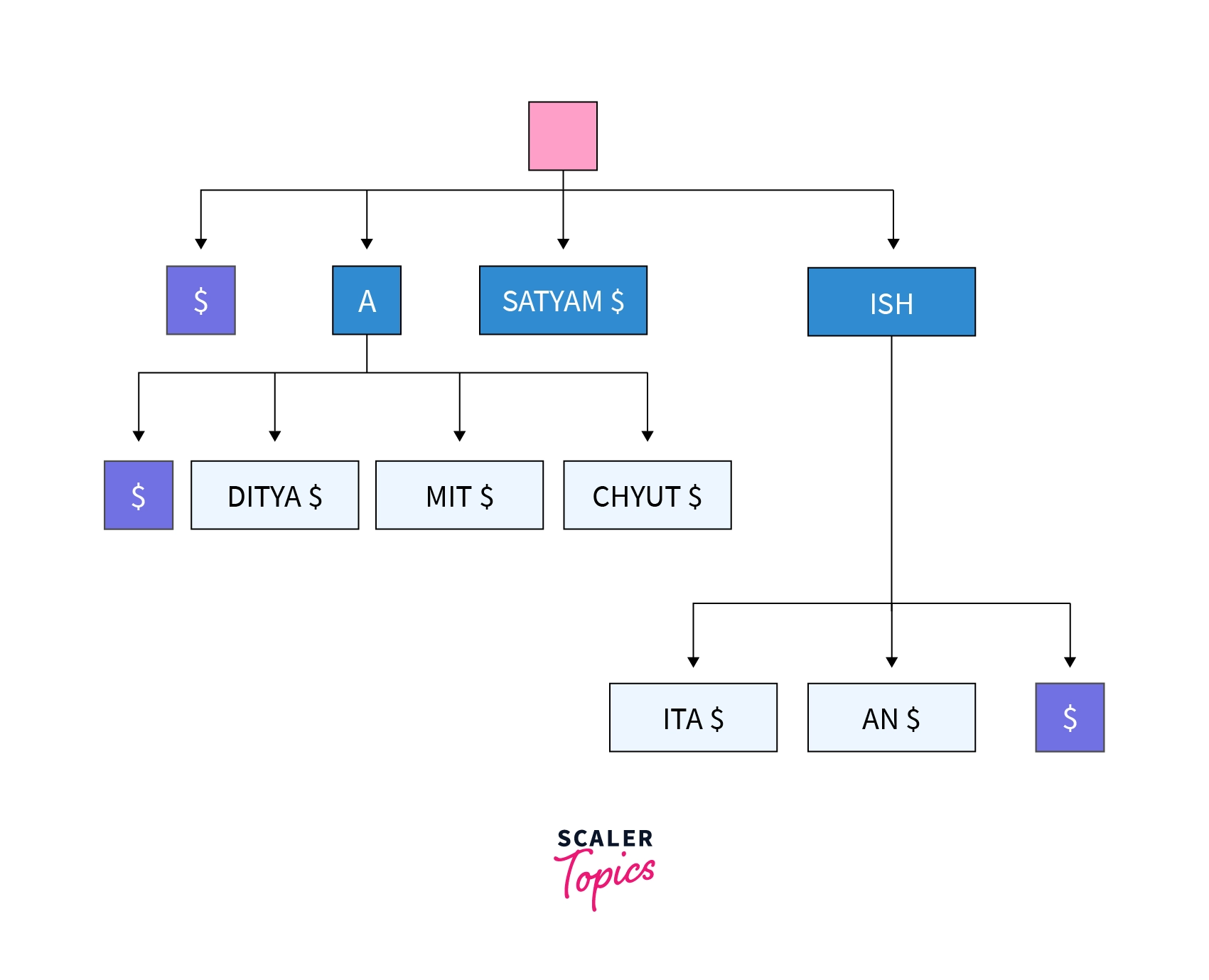
Here in terms of compression we are doing nothing but combining the further unique one nodes into a single node and adding '$' as a terminating character.
Functionality of Suffix Tree
A suffix tree for a string S of length n can be created in Theta (n) time if the letters come from an alphabet of integers with a polynomial range of -infinity to +infinity (in particular, this is true for fixed-sized alphabets). The majority of the time is spent sorting the letters into an O(n)-sized range for larger alphabets; on average, it takes O(nlog n) time. Imagine that over the string S of length n, a suffix tree has been constructed, then you can:
1. Look for strings:
- In O(m) time, determine whether a string P of length m is a substring.
- In O(m) time, find the very first occurrence of the sequences P1...PQ with a total length of m as substrings.
- In O(m+z) time, find all z occurrences of the patterns P1...PQ of length m in substrings.
2. Find properties of the strings.
- In Theta(n(i) + n(j) time, find the longest common substrings of the strings S(i) and S(j).
- In Theta(n + z) time, find all maximal pairings, maximal repeats, and supermaximal repeats.
- In Theta (n) time, find the longest repeated substrings.
- In Theta (n) time, find the most frequently occuring substrings of a minimum length.
- In Theta (n) time, find the shortest substrings that occur only once.
Applications of Suffix Tree
Suffix trees can also be used to handle a wide range of string challenges in applications, for example editing, free-text search, bioinformatics, and others. Among the most common uses are are listed below:
-
Exact String Matching- As previously stated, given the text's suffix tree, all occ occurrences of a pattern P may be discovered in O(P + occ). Even when the time spent building the suffix tree is factored in, this is asymptotically as fast as Knuth–Morris–Pratt for a single pattern for several patterns.
-
Computational Biology- Suffix trees are commonly used in Bioinformatics to find the repetitive patterns in a DNA strand. In a DNA strand, it can also be used to discover the longest common sub-string or sub-sequence. These methods are critical in the research of evolution and the identification of organismal commonalities.
-
Text Statistics- Every point in the suffix tree corresponds to a textual factor, and every factor corresponds to a point. As a result, despite the fact that the value obtained is typically Theta(n2), the number of different factors in the text equals the number of different point, which can be computed in O(n) time by traversing the suffix tree. The text's longest repeating factor is the longest string that appears at least two times in the text. It's represented by the suffix tree's most innermost internal node.
-
Longest Palindrome- A palindrome is just a sequence that is the opposite of itself. Racecar, for instance, is a palindrome. So this suffix tree concept is used to find the longest palindrome also.
-
Data compression- Data compression is the technique of encoding information with lesser bits than the original representation in signal processing.
Implementation of Suffix Tree
The complete tree can be expressed in Theta (n) space if each node and edge can be represented in Theta(1) space. The overall length of all strings on all the tree's edges is , but that each edge can be stored as the position and length of a substring of S, using Theta (n) computer words in total. A fibonacci word shows the worst-case space utilisation of a suffix tree, with all 2*n nodes.
The parent-child connections among nodes is a significant factor while creating a suffix tree. The most frequent method is to use sibling lists, which are connected lists. Each node has a pointer to its first child as well as the next node in the child list. Hash maps, sorted or unsorted arrays, and balanced search trees are some other solutions with effective running time attributes.
Now we're curious about:
- The cost of spotting a child on a particular character.
- The cost of a child's insertion.
- The cost of recruiting all of a node's children.
Let n be the alphabet's size. There are the costs listed below:
| Implement On | Extraction | Insertion | Traversal |
|---|---|---|---|
| Hash Map | Θ(1) | Θ(1) | O(n) |
| Sorted Arrays | O(logn) | O(n) | O(1) |
| Bitwise Sibling Tree | O(logn) | Θ(1) | Θ(1) |
| Balanced Search Tree | O(logn) | O(logn) | O(1) |
| Hash map + Sibling List | O(1) | O(1) | O(1) |
| Sibling Lists | O(n) | Θ(1) | Θ(1) |
Parallel Construction
Numerous parallel techniques for generating suffix trees have been presented.
A parallel algorithm for building suffix tree has recently been discovered with O(n) work (linear time) and span.On common multiple core platforms, the technique has strong scalability and can index about 3GB of the human genetic code whithin just 3 minutes that used a 40 core processor.
External construction
The space use of a suffix tree is considerably more than the real size of the sequence collections, regardless of the fact that it is linear. Creation of a huge document may require the use of removable storage techniques.
Theoretical studies for building suffix trees on removable storage exist. Farach-Colton, Ferragina, and Muthukrishnan (2000) proposed an algorithm that is conceptually optimum, with an I/O complexity equivalent to sorting. However, due to the algorithm's overall complexity, it has yet to be implemented in practise.
On either hand, practical works for developing disk-based suffix trees that expand to (a few) GB/hours have been published. TDD, TRELLIS, DiGeST and B2ST are some of the approaches.
Conclusion
- So after going through this article we have understood about that what actually suffix tree is along with an example with full explaination.
- We have gone through the full funtionality and implementation of the suffix tree along with it's applications.
- At last we have concluded with the parallel and external construction of suffix tree.
